Mauve
Mauve is a program to align multiple genomes.
Documentation: http://darlinglab.org/mauve/mauve.html
What it does:
- aligns genomes and identifies homologous blocks
- these are likely from a common ancestor or gained via horizontal transfer
- blocks may have moved or been inverted in the genome
Mauve - align three strains
We will align three genomes of Streptococcus pneumoniae.
Open Mauve.
- Go to
File :Align with Progressive Mauve
-
Add Sequence -
select the sequence(s). Use .fasta or .gbk files.
-
if using a reference sequence, add that first.
-
Align
-
Specify a name for the alignment.
-
Save
A console window will open and show the progress of the run.
When finished, the alignment will open:
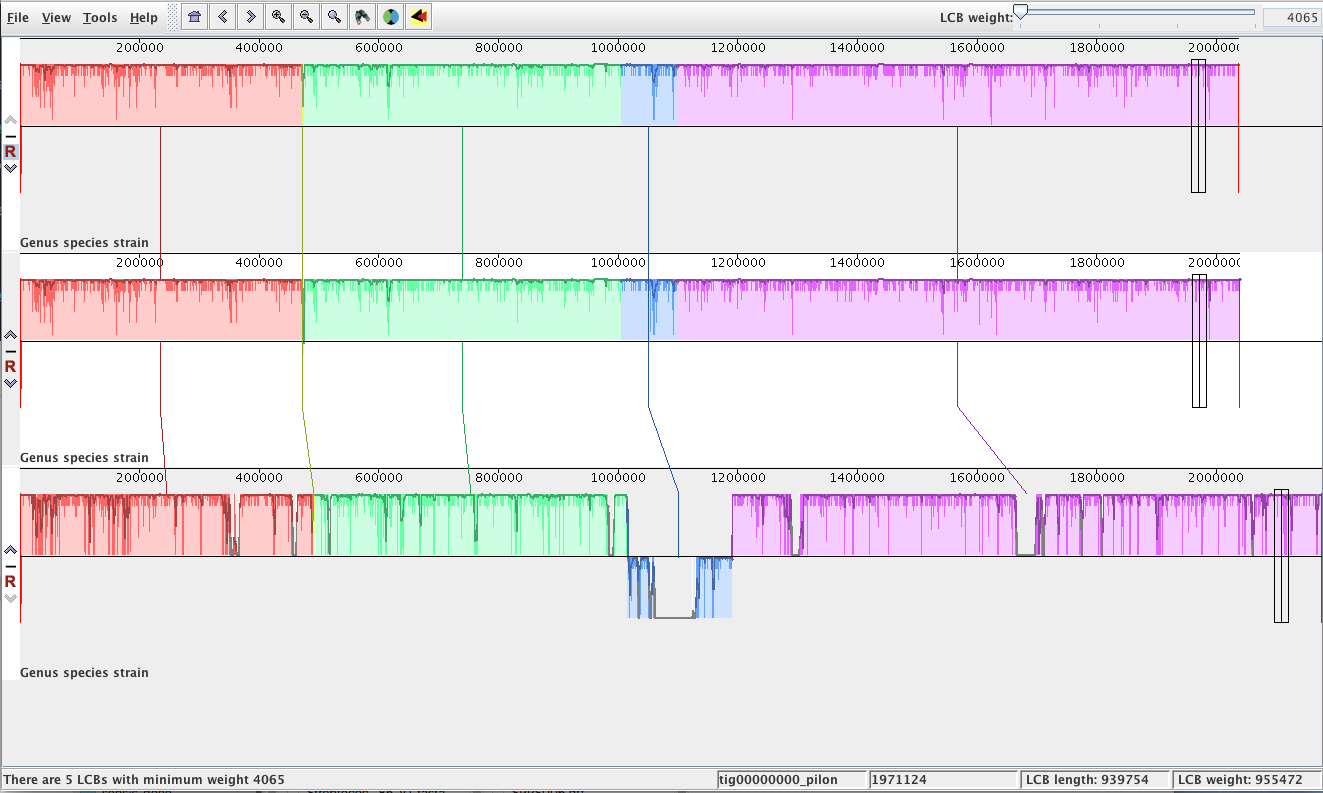
- Each row is a genome. Each coloured block is genetically similar.
- If you are using annotated genomes, zoom in (with the magnifying glass) to see annotations.
For a different view, go to
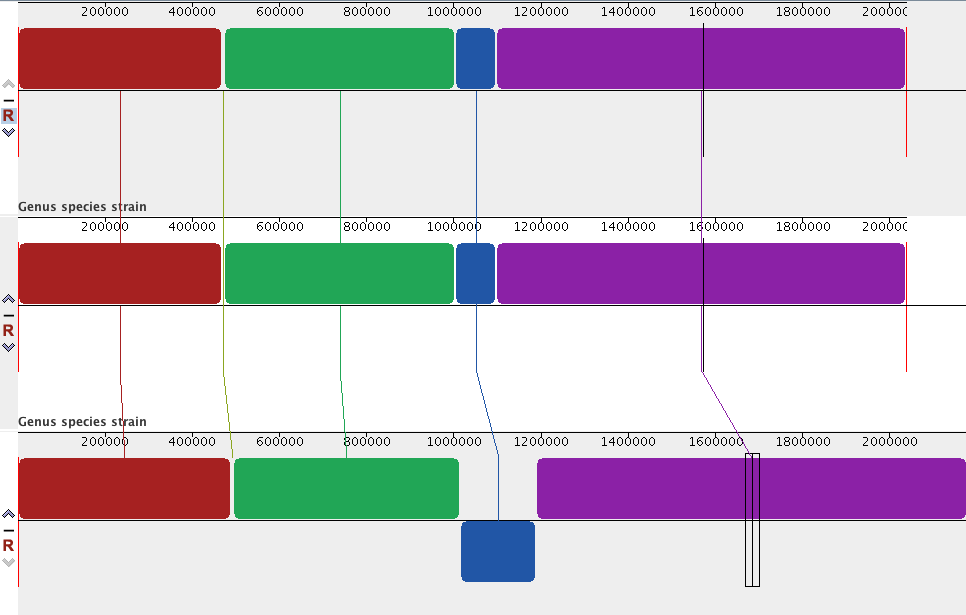

- Click on a block to align all genomes around that block.
- The blue block is inverted in genome 3 (i.e., the reverse complement).
Mauve - align two assemblies from the same sample
In this example, we will align two genomes from the same sample that have been assembled with different tools.
- Genome 1: Assembled from long reads; corrected with short reads.
- Genome 2: Assembled from short reads.
Align the genomes:
- Go to
File :Align with Progressive Mauve - Add sequences. Add the long-read assembly sequence first.
Align - Name
Save
View the alignment:
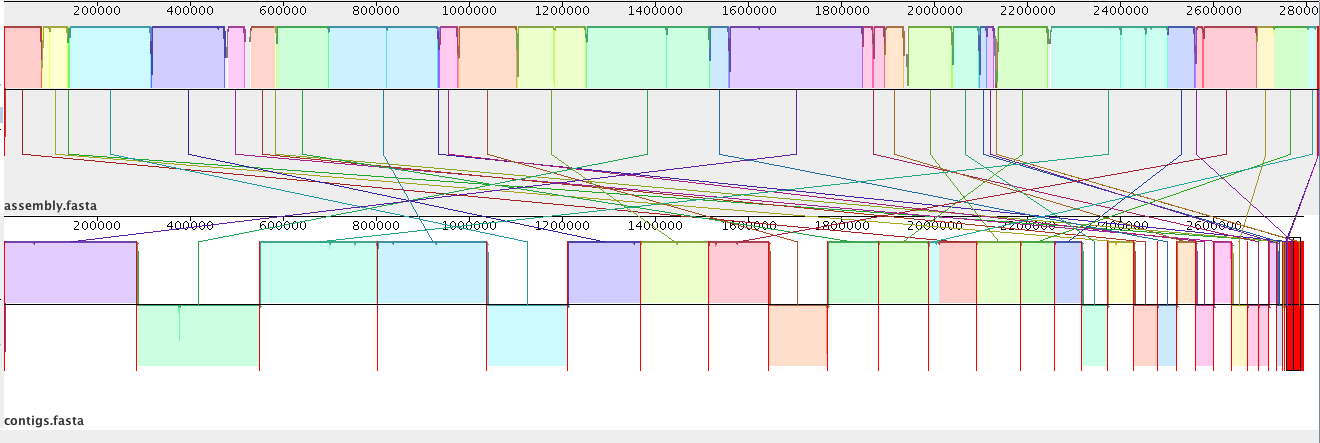
Genome 2 has many contigs as it has been assembled using short reads.
- These have been laid out in the order in which they appear in the file.
- We need to re-arrange these contigs to align with the reference genome (Genome 1).
Re-order the contigs in Genome 2:
- Go to
Tools :Move Contigs - Specify output folder
- Add sequences (add the long-read assembly first)
Start
The Mauve Console window will show the progress.
The re-ordered contigs will then be displayed:
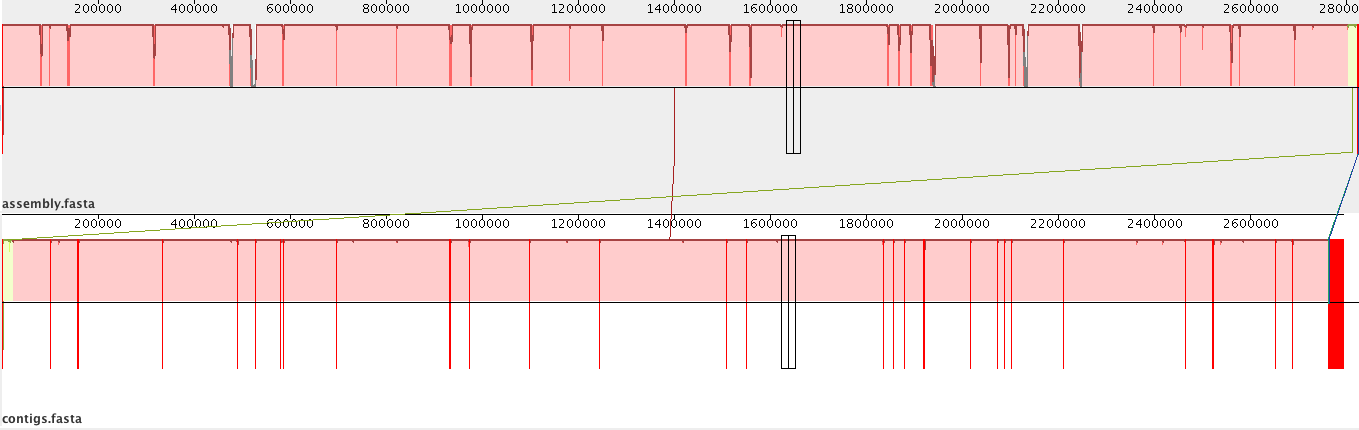
Most of the contigs in Genome 2 can be aligned to one (red) section of Genome 1.